Dataset overview
This study evaluates our method using three publicly available skeleton-based datasets: UI-PRMD37, KIMORE38, and EHE39. All three datasets provide two primary types of features: position and orientation of skeleton joints. These features are crucial for understanding the exercise performance of different populations, each presenting unique challenges in rehabilitation assessments. The position and orientation features are visualized in Fig. 3. An overview of the key characteristics of these datasets is provided in Table 1. The details of each dataset are described below.”
UI-PRMD Dataset contains 1326 exercise repetitions from 10 healthy subjects performing 10 rehabilitation exercises (e.g., side lunge, sit to stand, deep squat). Data was captured using Kinect v2 and Vicon motion capture. The datasets provide position and orientation features of skeleton joints. Subjects perform both correct and incorrect versions of exercises, simulating common errors in musculoskeletal rehabilitation. We use the Kinect v2 data for our experiments, following the consistent subset used in prior work to minimize errors.
KIMORE Dataset includes 2806 exercise repetitions from 78 subjects divided into three groups: Control Group – Experts (CG-E), Control Group – Non-Experts (CG-NE), and Group with Pain and Postural Disorders (GPP). Subjects perform five exercises (e.g., lifting arms, squatting) captured using the Kinect v2 sensor. The dataset includes RGB videos, depth videos, and skeleton joint positions and orientation information. It also includes clinical performance scores based on physician evaluations. This dataset offers a valuable benchmark for rehabilitation assessment, especially for subjects with motor dysfunctions.
EHE Dataset consists of 869 exercise repetitions from 25 elderly subjects performing six daily morning exercises (e.g., wave hands, hands up and down). Data was collected using Kinect v2 in a natural elderly home setting. The subjects had an average age of 68.4 years, with 10 diagnosed with Alzheimer’s disease (AD) at varying severity levels. The dataset provides position and orientation features of skeleton joints and is useful for analyzing exercise performance in elderly individuals, particularly those with AD.
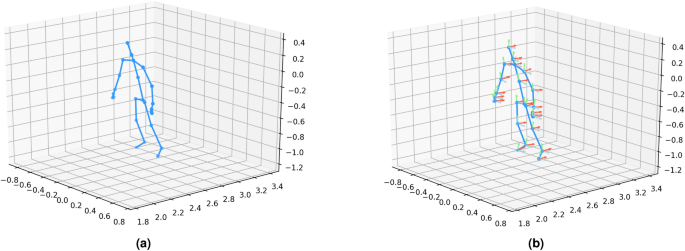
KIMORE dataset visualization.
Experimental setup
Data division strategies
To evaluate the generalization capability of the proposed models, we adopt two data split strategies: cross-subject division and random division. In the cross-subject division, the training and testing sets are constructed from disjoint subject identities—i.e., actions performed by one group of subjects are used for training, and actions from different subjects are used for testing. This simulates real-world deployment on previously unseen individuals. In the random division, all samples are randomly partitioned into training and testing sets without considering subject identities. This setting evaluates the average-case performance of the model. Both strategies are commonly used in skeleton-based action recognition and are designed to test model robustness under different generalization conditions.
Parameter settings
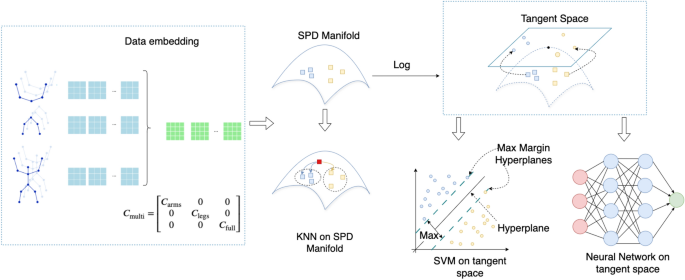
We evaluate the performance of three classification methods: KNN on the SPD manifold, SVM on the SPD manifold, and a Neural Network in Euclidean space. The KNN and Neural Network models are applied to two types of covariance matrix representations. The first is a full-body covariance matrix, computed using all skeleton joints. The second is a multi-scale covariance matrix, which includes separate covariance matrices for arms, legs, and full-body movements, later combined into a larger representation (as detailed in the Methodology section). Due to computational constraints, SVM is evaluated only with the full-body covariance matrix. We next describe the specific hyperparameters and configurations used for each of the three methods.
KNN classification on the SPD manifold parameters We assessed a range of distance metrics across multiple categories for KNN classification on the SPD manifold. Cholesky-based metrics included the Cholesky distance29 and the Log-Cholesky distance34. Log-Euclidean metrics comprised the Log-Euclidean distance30 and the Log-det distance35. Additionally, we evaluated the Affine-invariant distance31, the Euclidean distance, and information-theoretic metrics, such as the Kullback-Leibler divergence32 and its symmetric variant33. We selected \(k=3\) neighbors for KNN, as preliminary testing across \(k \in {1, 3, 5, 7}\) showed that \(k=3\) consistently yielded competitive performance under both the cross-subject and random division protocols, effectively balancing bias and variance.
Tangent space SPD SVM parameters For the SVM classification in the tangent space of the SPD manifold, we configured the model with the following hyperparameters: a regularization parameter \(C=0.1\), a training duration of 150 epochs, a learning rate of 0.01, and a batch size of 8. These values were selected based on standard practices in prior work and preliminary tuning on a validation set, balancing training stability and generalization. To optimize convergence and prevent overfitting, we employed an early stopping patience of 20 epochs and implemented a learning rate reduction strategy with a step factor of 5.0, reducing the learning rate until a minimum value of \(1 \times 10^{-4}\) was reached. We explored three distance metrics for mapping the SPD manifold to the tangent space: the affine-invariant metric, the Log-Euclidean metric, and the Euclidean distance, evaluating their impact on classification performance.
Neural Network-Based Method Parameters For our neural network-based approach, we adopted a network architecture detailed in the methodology section. The model was trained using the Adam optimizer with a learning rate of 0.001 and a batch size of 8 over 50 epochs. These values were chosen based on commonly used defaults in similar settings and verified to provide stable convergence during initial experiments. To regularize the optimization process, we applied a weight decay of \(1 \times 10^{-4}\) and configured the optimizer’s beta parameters to [0.9, 0.999], balancing the influence of past gradients and squared gradients for stable convergence.
Results and comparison
This section presents the classification results for KNN on the SPD manifold, Tangent Space SVM, and a neural network in Euclidean space, using two covariance matrix representations: full-body and multi-scale (arms, legs, full-body). Experiments were conducted on three datasets: UI-PRMD, KIMORE, and EHE. The following subsections present the results for each method in detail.
SPD manifold KNN with different distance metrics
To assess the impact of multi-scale covariance matrices, we evaluated the KNN classifier using various distance metrics. The results are presented in Table 2. The goal of this experiment is to determine whether the multi-scale representation enhances the distinctiveness of covariance matrices, thereby improving classification performance.
The findings reveal the following trends:
-
The multi-scale representation consistently outperforms the full-body representation across all datasets, except when using Euclidean-based metrics, where it shows no advantage. This suggests that the multi-scale approach improves the distinctiveness of covariance matrices for most metrics.
-
Cholesky29 and Kullback-Leibler divergence32 metrics achieve higher accuracy in 5 out of 6 cases when using the multi-scale representation, indicating these distance metrics benefit from the finer granularity of motion captured by the multi-scale matrices.
-
The symmetric variant of the Kullback-Leibler divergence33 metric improves accuracy in 4 out of 6 cases with the multi-scale representation, further reinforcing the advantages of this approach for capturing motion patterns.
-
Log-Euclidean30, Log-det35, and Affine-invariant31 metrics achieve higher accuracy in all 6 cases with multi-scale representation, suggesting that these metrics are highly compatible with the multi-scale approach.
-
The Euclidean-based metric does not benefit from multi-scale representation, likely due to the inherent mismatch between Euclidean geometry and the SPD manifold, where Euclidean distance fails to capture the complex structure of the covariance matrices.
These results confirm that the multi-scale covariance representation better captures fine-grained motion patterns across different body parts, enhancing distinctiveness and improving classification accuracy.
We also observed that the random split outperformed the cross-subject split across all datasets. This can be attributed to the nature of the KNN classifier, which relies on distance-based measurements. In the random split scenario, parts of the same subject’s data can end up in both the training and test sets, resulting in more similar data for both training and testing. This similarity improves KNN’s performance. In contrast, the subject cross-split ensures that training and testing data come from different subjects, which introduces more variation and makes the task more challenging, thus leading to lower performance.
KNN, while effective with certain distance metrics, has inherent limitations. It relies on predefined distance measures, struggles with high-dimensional data, and cannot model complex, non-linear relationships. These weaknesses make it less suitable for capturing the intricate patterns in multi-scale covariance matrices.
For the KNN classifier, we evaluated all three feature types—position (Pos), orientation (Ori), and their combination (PO)—and reported the averaged results across all action classes. Due to space constraints and the large number of configurations involved (3 datasets \(\times\) 8 distance metrics), we present only the Pos-based results in the main paper. Detailed modality-specific results (Pos, Ori, PO) under the Log-Euclidean metric, which serves as a representative choice due to its consistent performance across datasets, are provided in Supplementary Tables S1.
Tangent space linear SPD SVM under different metrics
To further analyze the effectiveness of different metrics on SPD matrix classification, we evaluated a Tangent Space Linear SPD SVM. Unlike KNN, which relies on local distance comparisons, SVM is a more powerful classifier that can find complex decision boundaries, particularly when operating in the tangent space of SPD manifolds. By mapping SPD matrices to a vector space, SVM can leverage linear separation techniques to improve classification accuracy.
Table 3 presents the classification accuracy for the UI-PRMD, KIMORE, and EHE datasets under cross-subject and random division protocols. The Log-Euclidean metric consistently achieved the highest accuracy across all settings, demonstrating its effectiveness in capturing the intrinsic structure of SPD matrices while maintaining computational tractability. In contrast, the AIRM, despite its theoretical appeal, produced the lowest performance in nearly all cases, possibly due to sensitivity to class imbalance or numerical instability. The Euclidean baseline, lacking geometric modeling, sometimes outperformed AIRM, particularly under random division, indicating that simpler approaches can occasionally be more robust.
These findings suggest that geometry-aware mappings, especially Log-Euclidean, generally enhance classification performance when paired with discriminative classifiers such as SVM. However, method selection should be guided by the data and application context, motivating further investigation into metric suitability and robustness.
We report SVM results using the multi-scale covariance representation with position (Pos) features, as its superiority has been clearly demonstrated in the KNN experiments. Due to the computational expense of training SVMs across multiple settings, we did not include separate evaluations using the single-scale representation and instead focused on the multi-scale variant, which offers better discriminative power.
For completeness, we provide additional SVM results using all three input types: multi-scale Pos, multi-scale Ori, and their combination multi-scale PO. These results, computed under the Log-Euclidean metric, are included in Supplementary Table S2.
Neural network approach in Euclidean space
While KNN and our Tangent Space Linear SPD SVM have shown effectiveness, they face inherent limitations. KNN relies on predefined distance metrics and struggles in high-dimensional spaces, while the Tangent Space Linear SPD SVM assumes that SPD matrices can be well-separated using a linear decision boundary. This assumption may not hold for complex SPD data, restricting its ability to capture intricate patterns. To address these challenges, we turn to neural networks, which can learn non-linear patterns, handle high-dimensional data more effectively, and automatically extract relevant features. This makes them a more robust alternative for classifying SPD matrices, leading to improved performance.
As demonstrated in Table 4, for the UI-PRMD dataset, the classification results across both cross-subject and random division protocols highlight the superior performance of the PO method (Position and Orientation-based information), which achieves the highest average accuracy of 92.40% in the cross-subject protocol and 96.15% in the random division protocol. The Ori method (Orientation-based information) ranks second, with 90.84% accuracy in the cross-subject protocol and 95.61% in the random division protocol. The Pos method (Position-based information) shows slightly lower performance at 86.12% in the cross-subject protocol. Our proposed methods consistently outperform the state-of-the-art methods MLE-O and MLE-PO, with Ori and PO achieving the best results. Notably, our methods surpass the MLE methods in 8 out of 10 exercises for the cross-subject protocol and in 7 out of 10 exercises for the random division protocol.
As demonstrated in Table 5, for the KIMORE dataset, our methods consistently outperform the previous state-of-the-art methods, MLE-PO and MLE-O, across both the cross-subject and random division protocols. Our approach achieves an average accuracy of 85.18% in the cross-subject protocol and 96.69% in the random division protocol, surpassing MLE-PO and MLE-O in both protocols. Specifically, under the cross-subject protocol, our methods outperform MLE-PO and MLE-O in 5 out of 8 exercises, while in the random division protocol, our methods achieve superior performance in all 8 exercises compared to MLE-PO and MLE-O.
As demonstrated in Table 6, for the EHE dataset, the classification results indicate that while the Pos method performs well, the PO method achieves the best overall performance, particularly in the random division protocol (95.79% average accuracy). In the cross-subject protocol, Pos leads with the highest average accuracy (87.59%), followed closely by MLE-PO and MLE-O, with MLE-O showing slightly lower accuracy than both Pos and MLE-PO. In the random division protocol, PO outperforms all other methods, achieving the best results in five out of six exercises. While Pos also performs well, it falls short of PO, with MLE-PO and MLE-O showing similar performance, and MLE-O trailing slightly behind.
In conclusion, across the UI-PRMD, KIMORE, and EHE datasets, our methods consistently outperform the state-of-the-art MLE-PO and MLE-O methods. The PO method, which combines position and orientation information, proves to be more robust and generally achieves the best results across both protocols in all three datasets. Even in cases where it doesn’t provide the absolute best accuracy, it still delivers strong performance. Overall, our approach demonstrates significant improvements over MLE-PO and MLE-O, confirming its effectiveness and superiority in human activity recognition.
Comparison with state-of-the-art
To evaluate the effectiveness of our proposed method, we compare it against several state-of-the-art approaches on the UI-PRMD, KIMORE, and EHE datasets using the cross-subject evaluation protocol. The results, presented in Table 7, demonstrate the superior performance of our approach in various settings.
GCN 25 leverage spatial dependencies in skeleton-based action recognition by applying graph convolutions to model joint interactions. This method achieved moderate accuracy across datasets, with performance varying between position-based (Pos) and orientation-based (Ori) features. AGCN 41 Adaptive GCNs (AGCN) enhance GCN by introducing adaptive graph structures that can dynamically learn relations between joints. While this method improves upon standard GCN, it struggles with orientation-based representations, particularly on the EHE dataset. MS-G3D 42 Multi-Scale Graph 3D Convolution (MS-G3D) incorporates multi-scale graph convolutions to capture both local and global motion patterns. This approach outperforms AGCN and GCN, particularly in orientation-based features, but still falls short in fully leveraging positional and orientation cues. CTR-GCN 43 Channel-wise Topology Refinement GCNs (CTR-GCN) introduce an adaptive channel-wise topology refinement strategy to better capture motion dynamics. This approach yields significant improvements over previous methods, particularly in the UI-PRMD and EHE datasets, demonstrating strong generalization capabilities. In contrast to these graph-based methods, SPDNet 44 learns directly from Symmetric Positive Definite (SPD) matrices using Riemannian geometry-based layers, allowing the network to operate on manifold-structured data while preserving its non-Euclidean properties.
MLE-O 40 employs ensemble learning in a two-stage process: first, training on joint positions for action recognition to identify key neurons, then refining the model for action correctness using orientation information, with pointwise weight multiplication ensuring HAR-relevant neurons influence correctness evaluation. This strategy enhances integration of spatial and orientation cues, improving performance over standard graph-based methods. MLE-PO 16 builds upon MLE-O by further refining the second learning stage, incorporating both position and orientation information for action correctness assessment. This enhancement allows the model to leverage spatial and orientation cues more effectively, leading to improved accuracy over its predecessor.
Our method significantly outperforms existing approaches across all datasets. In the position and orientation (Pos+Ori) category, our model achieves the highest accuracy of 92.40% on UI-PRMD and 85.05% on KIMORE, surpassing MLE-PO 16 in each case. For the EHE dataset, while MLE-PO achieves a higher accuracy (86.38%), our method achieves the best performance in the position (Pos) category with 87.59%, highlighting the robustness of our approach in integrating both spatial and orientation-based features and leading to improved generalization across datasets.
As we observe, the state-of-the-art methods for this task are GCN-based, as these methods are particularly well-suited for modeling the geometric structures inherent in human motion data. This makes them the natural choice for comparison, as they achieve state-of-the-art performance on tasks involving manifold-aware representations. While attention-based models, such as Transformers, have shown strong performance in other contexts, they operate in Euclidean space and do not account for the underlying SPD manifold geometry. As such, they are not directly comparable in this setting. Adapting attention-based models to non-Euclidean domains is a promising direction for future work but is beyond the scope of this study.
Computational cost comparison
In this subsection, we evaluate the computational efficiency of our proposed CNN-based model in comparison to state-of-the-art GCN-based methods on the UI-PRMD dataset. When measuring training time for one fold of 5-fold cross-validation on a Tesla V100-PCIE GPU, our model demonstrates superior efficiency. With a batch size of 8, the MLE-O40 model, a graph-based approach leveraging both position and orientation (Pos+Ori) features, requires approximately 77 seconds to train a single fold. In contrast, our CNN-based model, incorporating the same Pos+Ori features, completes the training in just 46 seconds under identical conditions, highlighting its computational advantage.
As shown in Table 8, our model has a significantly higher parameter count (e.g., 267.2M for Pos+Ori features compared to 3.1M–6.2M for GCN-based methods). This is primarily due to the dense fully connected layers in our Conv + MLP model, where each connection has a separate weight, leading to a quadratic increase in parameters with input size. In contrast, GCNs share parameters across nodes and typically avoid large fully connected layers, making them more parameter-efficient, especially for graph-structured data. Additionally, as noted in27, certain concepts or objects recur across various mathematical domains and often carry substantial implications. For example, a system involving three identical globes in robotics: originally represented in a nine-dimensional Euclidean space, can alternatively be modeled on a three-dimensional orthogonal group manifold. In our approach, we transform the data onto a SPD manifold to better capture its intrinsic geometric structure while also reducing computational complexity through fewer floating-point operations.
Despite the higher parameter count, our model achieves substantially lower FLOPs—0.6G compared to 1.9G–12.2G for GCN-based methods. This difference stems from the complexity of graph-based operations. In GCNs, each node aggregates features from its neighbors, which involves multiple matrix multiplications or neighbor-to-node feature propagation. This process becomes computationally expensive, particularly in large or dense graphs, where node degree and graph size both affect the cost. On the other hand, our Conv + MLP model operates on fixed-size local regions (e.g., image patches) with more localized operations, resulting in fewer FLOPs per layer despite having more parameters. The global aggregation in GCNs increases computational cost significantly, making them more FLOP-intensive, even with fewer parameters.
Ablation studies
The contributions of each model component, as evaluated on the UI-PRMD, KIMORE, and EHE datasets, are detailed in Table 9.
-
Baseline Model: Our full approach embeds skeleton data onto the SPD manifold, employs a multi-scale strategy (arms, legs, whole body) to form a combined covariance matrix, and applies a Log-Euclidean mapping to project the data onto the tangent space. This configuration achieves a mean accuracy of 86.28%.
-
Without Multi-Scale: Removing the multi-scale component reduces the mean accuracy to 85.50% (a drop of 0.78%), demonstrating the benefit of capturing localized dynamics.
-
Without SPD Embedding: Excluding the SPD embedding significantly degrades performance, lowering the mean accuracy to 81.95% (a drop of 4.33%), with a dramatic decline in the UI-PRMD position data (from 86.12% to 69.17%).
-
Without Log-Euclidean Mapping: Omitting the Log-Euclidean mapping results in a moderate decrease to 84.78% (a drop of 1.5%), confirming its importance in linearizing the manifold structure for effective classification.
These ablation studies clearly demonstrate that each module contributes to the overall performance of the model. The SPD embedding is particularly critical, as evidenced by the substantial drop in accuracy when it is removed. The multi-scale strategy and log-Euclidean mapping, while offering relatively smaller improvements individually, still play a significant role in refining the discriminative power of the learned representations. Collectively, these components synergize to enhance the network’s ability to evaluate rehabilitation exercise correctness effectively.
Discussion
General-purpose manifold learning methods, such as t-SNE, Isomap, and UMAP, are widely used for nonlinear dimensionality reduction but often fail to model the underlying geometry of the data, treating it as generic point clouds in Euclidean space. This can lead to suboptimal representations for data with specialized structures. To address this, our method leverages the Riemannian geometry of SPD manifolds, which is well-suited for modeling covariance matrices prevalent in motion and sensor-based data. Equipped with geometric tools like logarithmic and exponential maps, the SPD manifold preserves the intrinsic structure of second-order statistics, making it ideal for capturing the variability of rehabilitation movement patterns. Unlike other non-Euclidean spaces, such as Grassmannian manifolds (suited for linear subspaces) or hyperbolic manifolds (designed for hierarchical data), the SPD manifold provides a more precise and effective framework for this application45,46.
Although this study focuses on performance in terms of computational complexity and accuracy, real-world rehabilitation applications demand models that can run efficiently on resource-constrained devices. When using position features from the UI-PRMD dataset, the current model requires approximately 2086 MB of GPU memory during evaluation, as reported by nvidia-smi, suggesting it is feasible for deployment on modern edge devices with CUDA support. However, the SPD manifold representation in this work is based on complete action sequences, which may limit its responsiveness in real-time scenarios. A promising direction for future work is to incrementally accumulate the covariance matrix over shorter time windows, enabling early-stage classification and progressive evaluation as the movement unfolds. This would allow for more interactive, real-time feedback, which is particularly valuable in rehabilitation contexts.
In this work, the multi-scale covariance embedding is constructed using three intuitive partitions: arms, legs, and full body. This grouping reflects a straightforward anatomical structure, where limbs are naturally treated as coherent units in human motion. The choice was based on simplicity and interpretability, rather than dataset constraints or clinical guidelines. However, we acknowledge that this is just one of many possible partitioning strategies. Future work could explore alternative groupings, such as data-driven clustering or functional segmentation, to assess their potential impact on model performance and insight generation. A potential direction for future work is to analyze which joints or covariance components contribute most to the model’s decisions, in order to enhance transparency and support more informed use in rehabilitation settings.
We adopt the Log-Euclidean Riemannian metric to map SPD matrices to the tangent space, as it strikes a balance between mathematical rigor and computational tractability. Unlike AIRM—which offers desirable invariance properties but at high computational cost—the Log-Euclidean approach treats the space of SPD matrices as a Lie group under the matrix logarithm, resulting in a vector space structure that simplifies both implementation and optimization. Prior work has demonstrated its empirical effectiveness across various domains30, and we find it especially suitable for our setting, where fast and stable optimization is critical for learning from high-dimensional covariance descriptors.
In our Tangent Space Linear SVM experiments, the Log-Euclidean mapping consistently outperformed both AIRM and the Euclidean baseline. Although AIRM respects the manifold geometry more precisely, its use of the Riemannian mean as the base point may introduce bias in imbalanced datasets and lead to numerical instability. Interestingly, even on the balanced dataset, AIRM still underperformed, suggesting that other factors such as data variability or curvature sensitivity may also contribute. The Euclidean baseline, which ignores manifold geometry entirely, performed less effectively overall, further supporting the value of geometry-aware mappings in SPD classification.
The three classification pipelines explored in this study—KNN, tangent-space SVM, and neural network—are intentionally selected to reflect a range of modeling capacities and assumptions. KNN relies purely on manifold geometry and does not involve any training, SVM introduces supervised learning with linear decision boundaries in the tangent space, and the neural network approach enables hierarchical and nonlinear feature learning directly on the SPD manifold. While we do not fuse these models in this work, they provide complementary perspectives on how SPD representations support classification tasks, highlighting the manifold’s versatility across different learning paradigms. Future work could investigate hybrid strategies that combine these models, such as ensemble learning or multi-branch architectures, to further enhance robustness and generalization.
Beyond these three paradigms, other prototype-based strategies have also been developed. For example, Tang et al.47 proposed generalized learning Riemannian space quantization (GLRSQ), which learns discriminative prototypes directly on the SPD manifold using Riemannian distances. Compared with our Riemannian KNN classifier, which is instance-based and requires no training, GLRSQ produces a compact set of prototypes, yielding stronger generalization and more efficient inference once trained. A promising future direction is to combine these two paradigms—for instance, by initializing prototypes from nearest neighbors or refining them adaptively—thereby balancing the simplicity of non-parametric methods with the discriminative power of prototype learning.
link